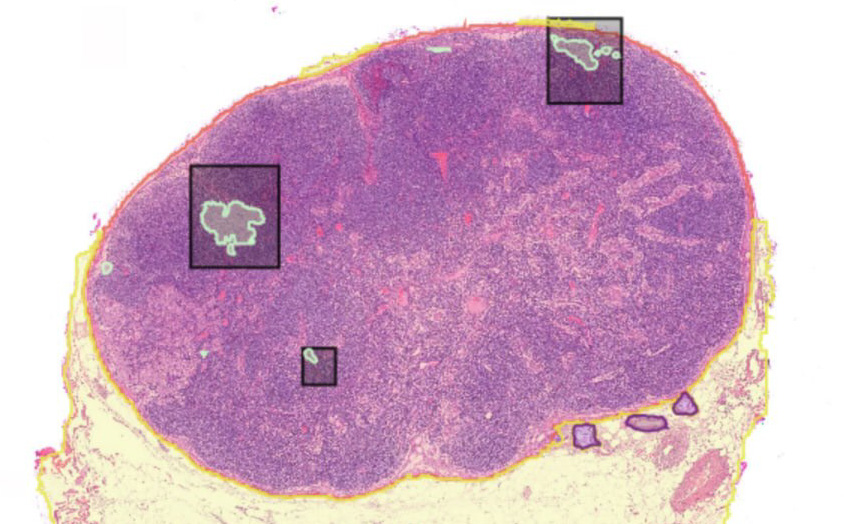
MetAssist is a digital pathology tool designed to reduce the burden of lymph node screening in surgical oncology. In colorectal cancer (CRC) staging, clinical guidelines require pathologists to examine at least 12 lymph nodes per resection specimen, with each node represented by one or more whole slide images (WSIs). Reviewing a dozen or more high-resolution slides per case is time-consuming and subject to sampling variability.
The project was initially developed for CRC [3] and is now being extended to multi-cancer lymph node screening. MetAssist combines Virchow2 [1], a large-scale pathology foundation model trained with self-supervised learning across mixed magnifications, with a MaskFormer decoder [2] for panoptic segmentation. This architecture jointly addresses two core tasks: segmentation of lymph node tissue regions within WSIs, and instance-level detection of metastatic foci within those regions. By automating the localization and triage of suspicious nodes, the tool assists pathologists in reaching accurate N-stage classifications faster and with greater consistency.
Key features include:
- Panoptic segmentation using Virchow2 [1] backbone with a MaskFormer decoder [2]
- Metastasis detection within segmented lymph node regions
- Metastatic size and shape estimation a key parameter in TNM staging criteria
- Physical lymph node count enabling automated N-stage determination
References
[1] Zimmermann, E., Vorontsov, E., Viret, J., Casson, A., Zelechowski, M., Shaikovski, G., … & Severson, K. (2024). Virchow2: Scaling self-supervised mixed magnification models in pathology. arXiv preprint arXiv:2408.00738.
[2] Cheng, B., Schwing, A., & Kirillov, A. (2021). Per-pixel classification is not all you need for semantic segmentation. Advances in Neural Information Processing Systems, 34, 17864-17875.
[3] Khan, A., Brouwer, N., Blank, A., Müller, F., Soldini, D., Noske, A., … & Zlobec, I. (2023). Computer-assisted diagnosis of lymph node metastases in colorectal cancers using transfer learning with an ensemble model. Modern Pathology, 36(5), 100118.